What We Do?

Natural Selection
Paragraph

Population Genetics
Paragraph

Bioinformatics
Paragraph
Evolutionary Genomics & Bioinformatics

Paragraph

Paragraph

Paragraph

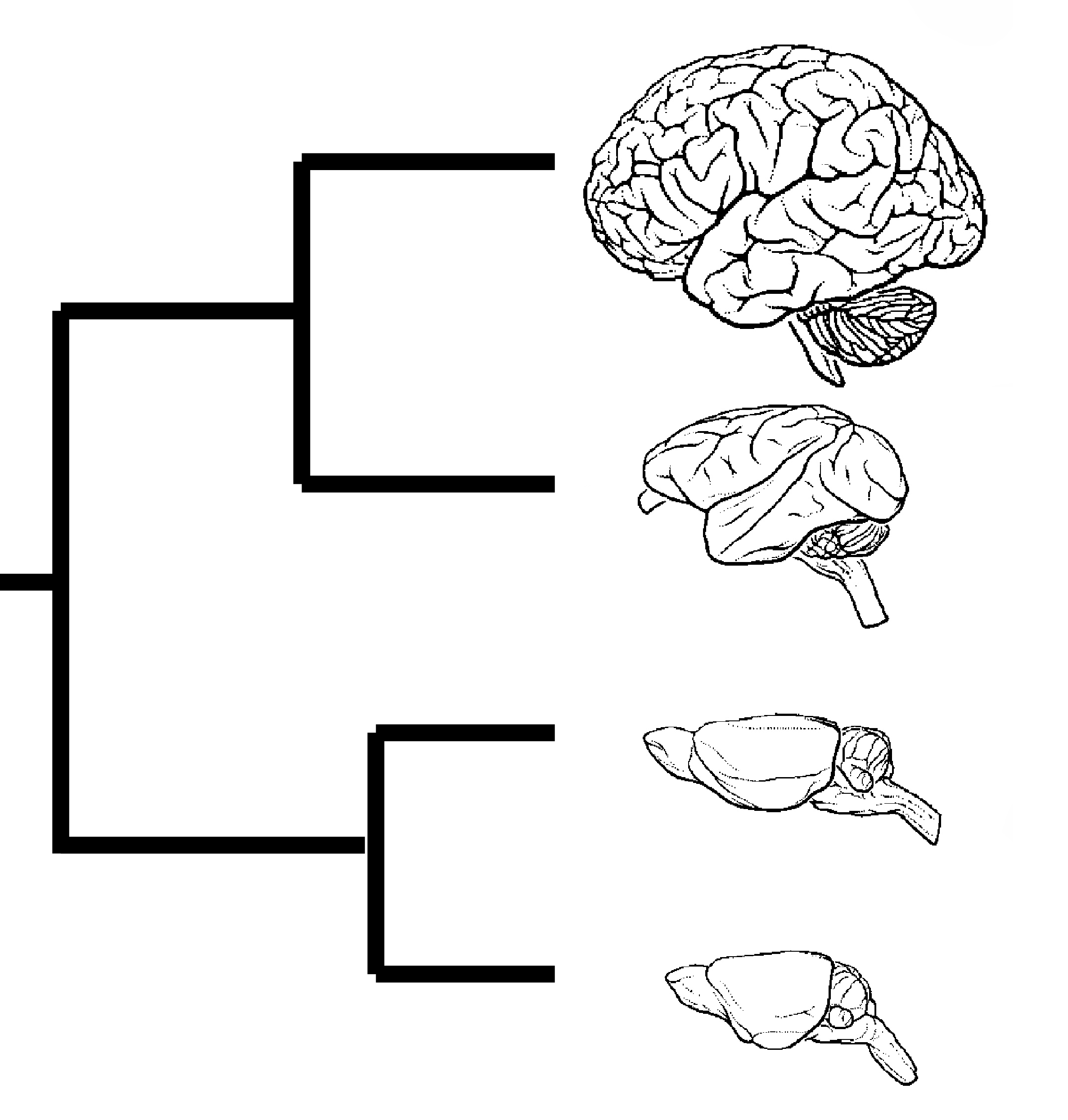 Accelerated evolution
Accelerated evolutionOver 5,400 different mammalian species exist globally and many have evolved highly distinctive phenotypic traits. The genomic mechanisms underlying these phenotypes are largely unknown but could be relevant for understanding human
disease. Accelerated genomic regions may have roles in shaping these important traits. Thus we develop new theories to study accelerated evolution driven by positive selection. Recently, we successfully identified the social enhancer
PAS1 that modulates social hierarchy in amniotes.

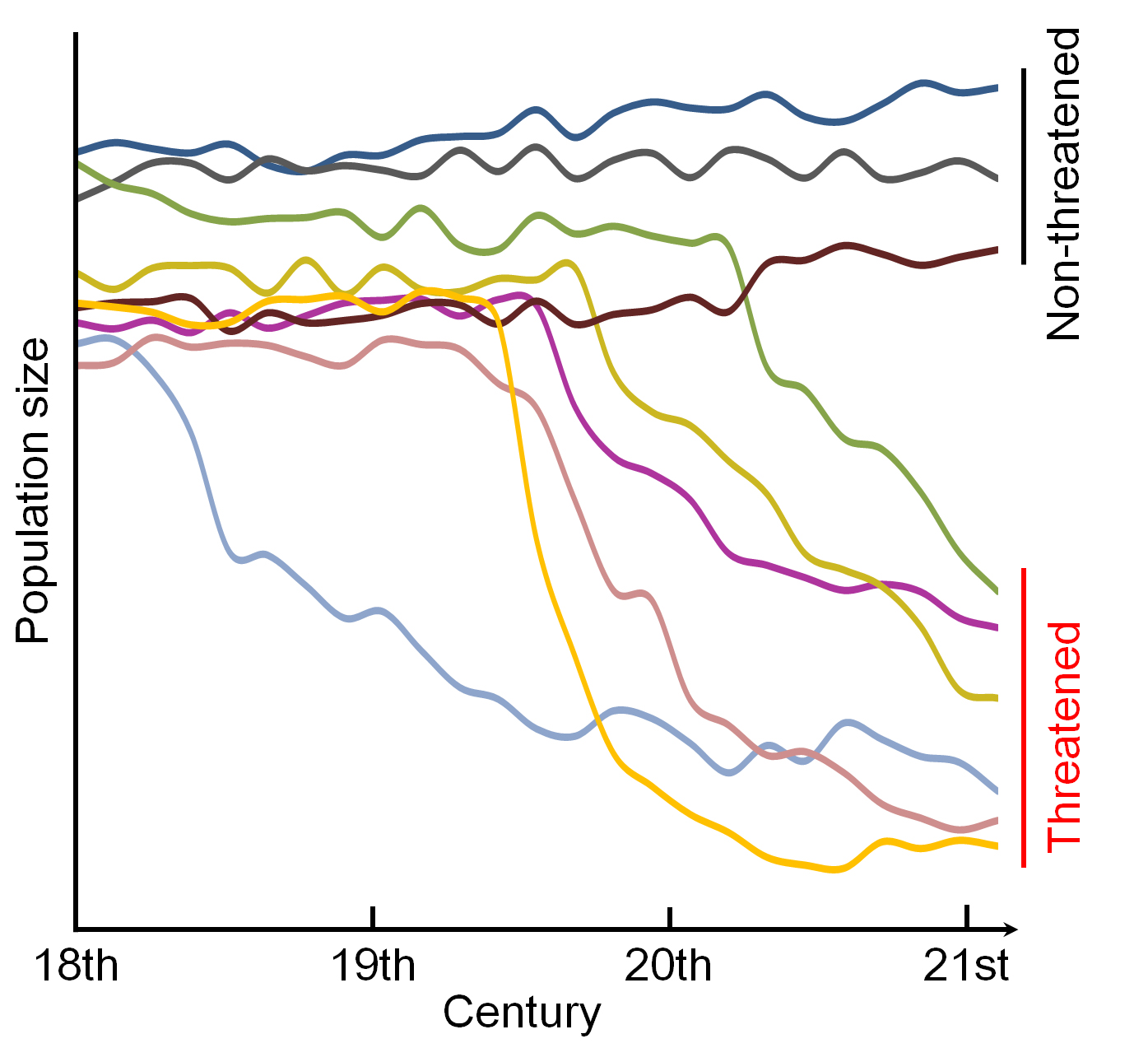
It can never been underestimated the importance of demography, for example, the demographic histories of human, chimpanzee, cat, fish, insects and plants. However, you cannot infer demography when there are no proper theories. Therefore,
by developing new theories, we aim to decipher demographic history of species. This is the beauty of theory as it can reveal evolutionary stories behind each species. These theories are essential for many research fields, for example,
precision medicine and conservation.
 Bioinformatics
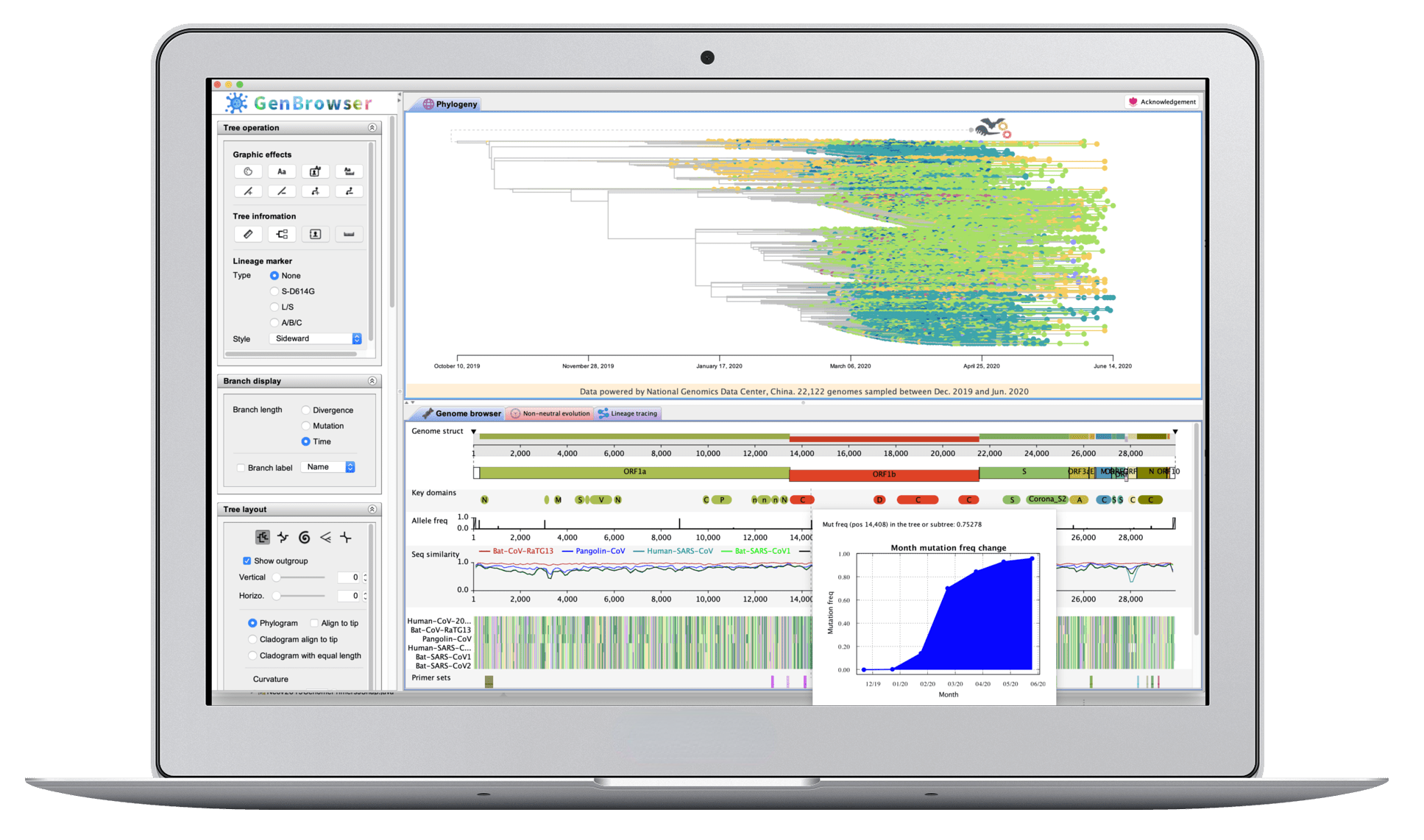
BioinformaticsMultiomic and evolutionary analyses have been booming recently, and interactive figures and software are believed to be important for these studies. We combine the advantages of cloud computing and desktop application, and design highly
interactive user interfaces and pipelines to bridge the gap between multiomic and evolutionary analyses. Moreover, COVID-19 is spreading across the world. To fight with SARS-CoV-2, the shared enemy of human community, we launch Coronavirus
GenBrowser for the public to rapidly access the latest information of SARS-CoV-2 genomes. It can closely monitor the frequency changes of mutations, identify the possible origin of newly emerged strains, and detect the non-neutral
evolution of SARS-CoV-2.

Natural selection is one of the important evolutionary forces. By detecting genes and non-coding regions subject to natural selection, it could help us to study the function of genes and non-coding regions, and the interactions between
genes and environment.
Our laboratory focuses on developing new theories for fast and efficient methods to detect positive selection. Moreover, since we introduced machine learning (or artificial intelligence) into the field
of population genetics ten years ago, we keep exploring possibilities by applying different machine learning methods, for example, deep convolutional neural networks, to detect positive selection. These studies are important for identifying
mutations associated with human metabolism and diseases.


 Psychobiotics
PsychobioticsPsychobiotics is a term used in preliminary research to refer to live bacteria that, when ingested in appropriate amounts, might confer a mental health benefit by affecting microbiota of
the host organism.
Recent advances in nutritional psychiatry point to the importance of the intestinal microbiota to human health and wellbeing. Amazingly, cognitive and emotional processes can be altered by microbes acting
through the brain-gut axis. The brain-gut axis provides bidirectional communication between the brain and the gut and includes the metabolically complex intestinal microbiota. However, we need new theories and methods to study why
and how gut bacteria can positively or negatively influence the mental process.